In eukaryotes, DNA is wrapped around histone octamers, which are composed of two copies each of histones H2A, H2B, H3 and H4. This structural arrangement of histones and DNA is known as chromatin. One of the main features of chromatin is that its structure can be altered to regulate the accessibility of the genome for diverse biological processes. The dynamic structure of chromatin is modulated mainly by the deposition of covalent modifications on DNA and histones, and the presence of histone variants. Different combinations of these chromatin “marks” throughout the genome are responsible for the epigenetic level of information that controls many cellular processes, including replication, transcription and DNA repair. Our lab is interested in understanding how chromatin participate in the cellular mechanisms responsible for maintaining genome and epigenome stability in multicellular eukaryotes. We are also interested in leveraging our expertise in chromatin biology to develop new tools and applications for plant genome engineering.
Regulation of chromatin replication by histone variants
In multicellular eukaryotes, the chromatin of proliferative cells typically contains three different histone H3 variants: H3.1, H3.3, and centromere-specific CenH3. While the protein sequence of CenH3 is considerably different from the other H3 variants, H3.1 and H3.3 are nearly identical in most eukaryotes, with only 4-5 amino acids varying between them. Despite being so similar, H3.1 and H3.3 had been hypothesized to play different cellular roles based on the fact that H3.1 is loaded on chromatin during replication to prevent nucleosome dilution, while H3.3 acts as a replacement histone H3 that is inserted at transcribed loci and other specific regions of the genome. These observations led to the “H3 barcode hypothesis” proposed by Hake and Allis in 2006, which stipulated that H3 variants can directly regulate chromatin function and contribute to epigenetic inheritance. Although this hypothesis was proposed more than 15 years ago, very few examples of H3-variant-specific functions have been demonstrated. Recently, we uncovered the first specific and widely conserved role for the H3.1 variant in regulating genome stability during replication in animals and plants. This finding substantiates the H3 barcode hypothesis and opens up many new and exciting research directions regarding the roles of the H3.1 variant in DNA repair and genome stability.
Our interest in the H3.1 variant is linked to our long-term research of the interplay between chromatin and genome stability using the flowering plant Arabidopsis thaliana as a model system. Previously, we discovered that the enzymes ATXR5 and ATXR6 (ATXR5/6) specifically mono-methylate H3.1 at lysine 27 (H3.1K27me1) during DNA replication. In atxr5/6 double mutants, H3.1K27me1 levels are depleted, which induces several phenotypes, including transposon de-repression, chromatin decondensation, and most importantly, genomic amplification of heterochromatic sequences. We recently discovered that H3.1K27me1 is responsible for inhibiting the activity of the conserved DNA repair protein TONSOKU (TSK; known as TONSL in animals), which was shown to resolve stalled or broken replication forks via homologous-recombination-mediated repair in mammals. TSK/TONSL contains a tetratricopeptide repeat (TPR) domain that reads alanine 31 (A31) of H3.1, which is replaced by serine or threonine in the H3.3 variant. This allows TSK/TONSL to specifically interact with soluble H3.1/H4 tetramers and be deposited on chromatin via the replication-dependent CAF1-mediated histone deposition pathway. The interaction of TSK/TONSL with H3.1 is proposed to be responsible for localizing these proteins at sites of DNA replication, which facilitates their activity in resolving stalled or broken replication forks (Fig. 1).
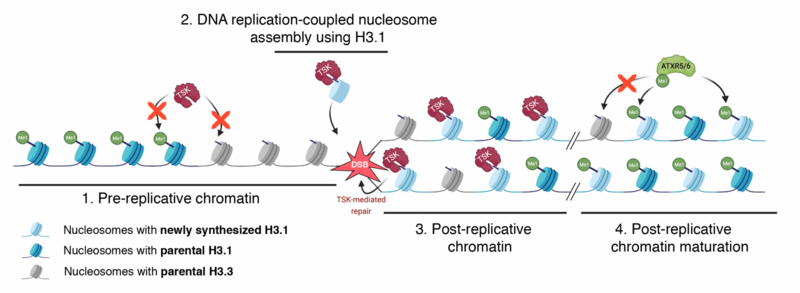
Figure 1: Model depicting the interplay between H3.1, TSK, and ATXR5/6 during replication. Step 1. TSK cannot interact with chromatin containing H3.3K27me0 or H3.1K27me1. Step 2. Newly synthesized H3.1 (H3.1K27me0) in complex with TSK are inserted at replication forks. Step 3. DSBs caused by broken replication forks are repaired by TSK. Step 4. Mono-methylation of newly inserted H3.1 (but not H3.3) at K27 by ATXR5/6 prevents binding of TSK.
Over the next few years, our lab is planning to study the molecular roles played TSK/TONSL in mediating genomic and epigenomic stability during DNA replication. We are also interested in uncovering new functions for histone H3 variants in chromatin. This NIH-funded work should have an impact in identifying mechanisms in human cells that serve to prevent genetic and/or epigenetic mutations that may arise during replication and be responsible for diseases like cancer.
Genome engineering and chromatin
Basic plant biology research and crop improvement rely on our ability to induce site-specific genetic alterations. CRISPR/Cas9 and related technologies have been shown to be powerful methods to induce targeted mutations in many eukaryotes, including plants. Although bacterial-derived CRISPR systems are efficient in producing edits, they can still benefit from optimization for use in the chromatin-based environment found in eukaryotes. Similarly to other processes like transcription, replication and DNA repair, accessibility to DNA is critical for efficient CRISPR/Cas9-induced modifications. This suggests that some genomic regions (e.g., heterochromatin) will be more challenging to edit with CRISPR/Cas9 than others, and this could be particularly important in plants. Most crops are flowering plants and genome size, complexity and organization vary greatly in this clade due to expansion of transposons and other repetitive sequences (e.g., ~85% of the maize genome consists of transposons). Also, many crops are polyploid and thus require multiple editing events to alter a single gene, a situation that would also benefit from increased editing efficiency of CRISPR/Cas9. For all these reasons and others, our lab is interested in optimizing gene editing systems for use in plants. We used our knowledge of chromatin biology and pathways regulating genome stability (e.g., DNA repair pathways) to improve plant genome engineering, design new applications in plants for gene editing, and solve current issues in agriculture. For example, we demonstrated a few years ago that exposing plants to heat stress resulted in an increase of CRISPR/Cas9-induced mutations in Arabidopsis and Citrus plants (Fig. 2). Optimizing editing in crops like Citrus cultivars (e.g., orange and lemon trees) is important, as it may provide new genetic variants in the future that are resistant to diseases like Huanglongbing, which is currrently decimating the U.S. Citrus industry.

Our lab currently has projects related to increasing the efficiency of gene targeting (i.e., precise editing that relies on homologous recombination-based DNA repair) in plants. We are also interested in optimizing multiplex CRISPR/Cas9 systems to perform large-scale engineering of plant genomes.
Figure 2: Effect of heat stress on gene editing by CRISPR/Cas9 in Citrus plants (Carrizo citrange). Young Citrus seedlings expressing Cas9 and a sgRNA targeting the PDS gene of Citrus. pds mutations in plants induce the presence of white tissues. In this gene editing experiment, leaf tissues formed before exposing the plants to heat stress are mostly green, while leaf tissues formed after exposing plants to heat stress are white. This demonstrates the impact of heat stress in increasing the efficiency of CRISPR/Cas9 editing.